Journal archives for December 2017
December 18, 2017

December 22, 2017
Fish Taxonomy
I added a script here that compares Fishbase to the 6 fish classes on iNaturalist (ie ray-finned fishes + sharks&rays + hagfish etc.). Ignoring the following 'explicit discrepancies' iNat now completely matches Fishbase. (one caveat is that the script uses a Fishbase API here https://fishbase.ropensci.org/ which I guess uses database copy from http://www.fishbase.org/ and has a few differences from the website). The 'explicit discrepancies' are as follows:
discrepancies = [
#these arn't in the Fishbase API, but are in the Fishbase website so leaving them
{fishbase: [], inat: ["Neomyxine caesiovitta"]},
{fishbase: [], inat: ["Chimaera carophila"]},
{fishbase: [], inat: ["Hypanus dipterurus"]},
{fishbase: [], inat: ["Neotrygon trigonoides"]},
{fishbase: [], inat: ["Neotrygon australiae"]},
{fishbase: [], inat: ["Potamotrygon rex"]},
{fishbase: [], inat: ["Bathytoshia lata"]},
{fishbase: [], inat: ["Etmopterus lailae"]},
{fishbase: [], inat: ["Eptatretus cryptus"]},
{fishbase: [], inat: ["Eptatretus poicilus"]},
{fishbase: [], inat: ["Micropterus haiaka"]},
{fishbase: [], inat: ["Stegastes marginatus"]},
{fishbase: [], inat: ["Pomacentrus maafu"]},
{fishbase: [], inat: ["Sirembo amaculata"]},
{fishbase: [], inat: ["Sirembo wami"]},
{fishbase: [], inat: ["Pomacentrus magniseptus"]},
{fishbase: [], inat: ["Pomacentrus micronesicus"]},
{fishbase: [], inat: ["Altrichthys alelia"]},
{fishbase: [], inat: ["Prognathodes basabei"]},
{fishbase: [], inat: ["Osteomugil engeli"]},
#iSeahorse discrepancies
#new species in iseahorse not in fishbase
{fishbase: [], inat: ["Hippocampus casscsio"]},
#lumped in iseahorse
{fishbase: ["Hippocampus multispinus","Hippocampus hendriki","Hippocampus grandiceps","Hippocampus angustus"], inat: ["Hippocampus angustus"]},
{fishbase: ["Hippocampus montebelloensis","Hippocampus zebra"], inat: ["Hippocampus zebra"]},
{fishbase: ["Hippocampus semispinosus","Hippocampus queenslandicus","Hippocampus spinosissimus"], inat: ["Hippocampus spinosissimus"]},
{fishbase: ["Hippocampus suezensis","Hippocampus kelloggi"], inat: ["Hippocampus kelloggi"]},
{fishbase: ["Hippocampus lichtensteinii", "Hippocampus zosterae"], inat: ["Hippocampus zosterae"]},
{fishbase: ["Hippocampus curvicuspis","Hippocampus histrix"], inat: ["Hippocampus histrix"]},
#Will lump these once we have range maps from iSeahorse
###{fishbase: ["Hippocampus alatus", "Hippocampus spinosissimus"], inat: ["Hippocampus spinosissimus"]},
###{fishbase: ["Hippocampus biocellatus","Hippocampus trimaculatus"], inat: ["Hippocampus trimaculatus", "Hippocampus planifrons", "Hippocampus dahli"]},
###{fishbase: ["Hippocampus procerus","Hippocampus whitei"], inat: ["Hippocampus whitei"]},
###{fishbase: ["Hippocampus waleananus","Hippocampus satomiae"], inat: ["Hippocampus satomiae"]},
###{fishbase: ["Hippocampus fuscus","Hippocampus borboniensis","Hippocampus kuda"], inat: ["Hippocampus kuda"]},
#Explicit deviations for Fishbase recommended by maractwin et al
{fishbase: ["Cheilinus fasciatus"], inat: ["Cheilinus quinquecinctus","Cheilinus fasciatus"]},
{fishbase: ["Chrysiptera brownriggii"], inat: ["Chrysiptera leucopoma","Chrysiptera brownriggii"]},
{fishbase: ["Poecilia sphenops"], inat: ["Poecilia thermalis","Poecilia sphenops"]},
{fishbase: ["Synodus variegatus"], inat: ["Synodus houlti", "Synodus variegatus"]},
{fishbase: ["Antennatus coccineus"], inat: ["Antennarius nummifer","Antennatus coccineus"]},
{fishbase: ["Synodus myops"], inat: ["Trachinocephalus myops", "Trachinocephalus trachinus"]},
{fishbase: ["Centrolabrus melanocercus"], inat: ["Symphodus melanocercus"]},
{fishbase: ["Nosferatu pantostictus"], inat: ["Herichthys pantostictus"]},
{fishbase: ["Paraneetroplus bifasciatus"], inat: ["Paraneetroplus bifasciatus","Vieja bifasciata"]},
{fishbase: ["Paraneetroplus fenestratus"], inat: ["Vieja fenestrata"]},
{fishbase: ["Barbodes semifasciolatus"], inat: ["Puntius semifasciolatus"]},
{fishbase: ["Leucos aula"], inat: ["Rutilus aula"]},
{fishbase: ["Ericymba buccata"], inat: ["Notropis buccatus"]},
{fishbase: ["Lethrinus lentjan"], inat: ["Lethrinus punctulatus","Lethrinus lentjan"]},
{fishbase: ["Pagrus auratus"], inat: ["Pagrus auratus","Chrysophrys auratus"]},
#Mystery synonyms... Need to vet and keep of find destinations for/inactivate these names not in Fishbase
{fishbase: [], inat: ["Tosanoides obama"]},
{fishbase: [], inat: ["Suttonia kermadecensis"]},
{fishbase: [], inat: ["Suttonia divaricata"]},
{fishbase: [], inat: ["Amblyeleotris gymnocephalus"]},
{fishbase: [], inat: ["Trimma finistrinum"]},
{fishbase: [], inat: ["Rhinogobius nganfoensis"]},
{fishbase: [], inat: ["Rhinogobius vinhensis"]},
{fishbase: [], inat: ["Cryptocentroides argulus"]},
{fishbase: [], inat: ["Pseudogobius melanosticta"]},
{fishbase: [], inat: ["Gobiopsis liolepis"]},
{fishbase: [], inat: ["Oxyurichthys zeta"]},
{fishbase: [], inat: ["Dotsugobius bleekeri"]},
{fishbase: [], inat: ["Hazeus diacanthus"]},
{fishbase: [], inat: ["Gobioides broussonetii"]},
{fishbase: [], inat: ["Egglestonichthys ulbubunitj"]},
{fishbase: [], inat: ["Schismatogobius insignis"]},
{fishbase: [], inat: ["Omobranchus fasciolaticeps"]},
{fishbase: [], inat: ["Pseudocaranx georgianus"]},
{fishbase: [], inat: ["Pseudocrenilabrus pyrrhocaudalis"]},
{fishbase: [], inat: ["Thorichthys maculipinnis"]},
{fishbase: [], inat: ["Cichlasoma nebuliferus"]},
{fishbase: [], inat: ["Vieja bimaculata"]},
{fishbase: [], inat: ["Australoheros autochthon"]},
{fishbase: [], inat: ["Rocio spinosissima"]},
{fishbase: [], inat: ["Cribroheros robertsoni"]},
{fishbase: [], inat: ["Cribroheros longimanus"]},
{fishbase: [], inat: ["Cribroheros alfari"]},
{fishbase: [], inat: ["Cribroheros rostratus"]},
{fishbase: [], inat: ["Cribroheros bussingi"]},
{fishbase: [], inat: ["Cribroheros diquis"]},
{fishbase: [], inat: ["Cribroheros altifrons"]},
{fishbase: [], inat: ["Cribroheros rhytisma"]},
{fishbase: [], inat: ["Kihnichthys ufermanni"]},
{fishbase: [], inat: ["Rheoheros lentiginosus"]},
{fishbase: [], inat: ["Darienheros calobrensis"]},
{fishbase: [], inat: ["Maskaheros argenteus"]},
{fishbase: [], inat: ["Maskaheros regani"]},
{fishbase: [], inat: ["Teleocichla preta"]},
{fishbase: [], inat: ["Oscura heterospila"]},
{fishbase: [], inat: ["Rheoheros coeruleus"]},
{fishbase: [], inat: ["Plectorhinchus caeruleonothus"]},
{fishbase: [], inat: ["Percina apina"]},
{fishbase: [], inat: ["Parupeneus heptacantha"]},
{fishbase: [], inat: ["Pomadasys approximans"]},
{fishbase: [], inat: ["Mullus barbatus"]},
{fishbase: [], inat: ["Apogon infuscus"]},
{fishbase: [], inat: ["Pristicon trimaculata"]},
{fishbase: [], inat: ["Sphaeramia nematopterus"]},
{fishbase: [], inat: ["Foa yamba"]},
{fishbase: [], inat: ["Jaydia carinata"]},
{fishbase: [], inat: ["Ozichthys albimaculosus"]},
{fishbase: [], inat: ["Jaydia truncatus"]},
{fishbase: [], inat: ["Diplodus sargus"]},
{fishbase: [], inat: ["Dentex carpenteri"]},
{fishbase: [], inat: ["Crenidens macracanthus"]},
{fishbase: [], inat: ["Girella fremenvillei"]},
{fishbase: [], inat: ["Scorpis hectori"]},
{fishbase: [], inat: ["Scorpis boops"]},
{fishbase: [], inat: ["Scorpis australis"]},
{fishbase: [], inat: ["Monotaxis heterodon"]},
{fishbase: [], inat: ["Nemadactylus macroptera"]},
{fishbase: [], inat: ["Parascolopsis rufomaculata"]},
{fishbase: [], inat: ["Nemadactylus carponotatus"]},
{fishbase: [], inat: ["Nemadactylus concinnus"]},
{fishbase: [], inat: ["Cheilodactylus antonii"]},
{fishbase: [], inat: ["Cheilodactylus aspersus"]},
{fishbase: [], inat: ["Cheilodactylus carmichaelis"]},
{fishbase: [], inat: ["Microdesmus longispinnis"]},
{fishbase: [], inat: ["Gobiomorphus gobiodes"]},
{fishbase: [], inat: ["Ditrema temminckii"]},
{fishbase: [], inat: ["Boroda malua"]},
{fishbase: [], inat: ["Eucinostomus californiensis"]},
{fishbase: [], inat: ["Pseudocalliurichthys goodladi"]},
{fishbase: [], inat: ["Bathycallionymus bifilum"]},
{fishbase: [], inat: ["Bathycallionymus kailolae"]},
{fishbase: [], inat: ["Calliurichthys afilum"]},
{fishbase: [], inat: ["Calliurichthys australis"]},
{fishbase: [], inat: ["Calliurichthys grossi"]},
{fishbase: [], inat: ["Calliurichthys ogilbyi"]},
{fishbase: [], inat: ["Foetorepus apricus"]},
{fishbase: [], inat: ["Foetorepus grandoculis"]},
{fishbase: [], inat: ["Pterosynchiropus occidentalis"]},
{fishbase: [], inat: ["Repomucenus filamentosus"]},
{fishbase: [], inat: ["Repomucenus keeleyi"]},
{fishbase: [], inat: ["Repomucenus meridionalis"]},
{fishbase: [], inat: ["Repomucenus sphinx"]},
{fishbase: [], inat: ["Repomucenus sublaevis"]},
{fishbase: [], inat: ["Repomucenus belcheri"]},
{fishbase: [], inat: ["Stichaeus punctatus"]},
{fishbase: [], inat: ["Heteropriacanthus carolinus"]},
{fishbase: [], inat: ["Heteropriacanthus fulgens"]},
{fishbase: [], inat: ["Monodactylus falciformes"]},
{fishbase: [], inat: ["Olisthops brownii"]},
{fishbase: [], inat: ["Sillago burra"]},
{fishbase: [], inat: ["Uranoscopus terraereginae"]},
{fishbase: [], inat: ["Hypopterus macroptera"]},
{fishbase: [], inat: ["Aurigequula longispinis"]},
{fishbase: [], inat: ["Psenes hillii"]},
{fishbase: [], inat: ["Emmelichthys nitidus"]},
{fishbase: [], inat: ["Matsubaraea fusiformis"]},
{fishbase: [], inat: ["Barbus lepineyi"]},
{fishbase: [], inat: ["Barbus moulouyensis"]},
{fishbase: [], inat: ["Barbus pallaryi"]},
{fishbase: [], inat: ["Barbus setivimensis"]},
{fishbase: [], inat: ["Puntius euspilurus"]},
{fishbase: [], inat: ["Dionda flavipinnis"]},
{fishbase: [], inat: ["Scaphesthes tamusuiensis"]},
{fishbase: [], inat: ["Anabarilius liui"]},
{fishbase: [], inat: ["Squalidus gracilis"]},
{fishbase: [], inat: ["Squalidus japonicus"]},
{fishbase: [], inat: ["Sarcocheilichthys variegatus"]},
{fishbase: [], inat: ["Physoschistura chulabhornae"]},
{fishbase: [], inat: ["Orthrias angorae"]},
{fishbase: [], inat: ["Beaufortia schaueri"]},
{fishbase: [], inat: ["Beaufortia orbifolia"]},
{fishbase: [], inat: ["Beaufortia micrantha"]},
{fishbase: [], inat: ["Meuschenia scabra"]},
{fishbase: [], inat: ["Aluterus abassai"]},
{fishbase: [], inat: ["Mola tecta"]},
{fishbase: [], inat: ["Pterois paucispinula"]},
{fishbase: [], inat: ["Acanthostracion bucephalus"]},
{fishbase: [], inat: ["Chilomycterus spinosus"]},
{fishbase: [], inat: ["Scorpaenopsis insperata"]},
{fishbase: [], inat: ["Scorpaena africana"]},
{fishbase: [], inat: ["Scorpaena aculeata"]},
{fishbase: [], inat: ["Sebastapistes mauritianus"]},
{fishbase: [], inat: ["Sebastolobus varispinis"]},
{fishbase: [], inat: ["Trachyscorpia cristulata"]},
{fishbase: [], inat: ["Onigocia macrocephala"]},
{fishbase: [], inat: ["Platycephalus angustus"]},
{fishbase: [], inat: ["Platycephalus australis"]},
{fishbase: [], inat: ["Liparis madrensis"]},
{fishbase: [], inat: ["Liparis makinoana"]},
{fishbase: [], inat: ["Notoliparis stewarti"]},
{fishbase: [], inat: ["Paraliparis copei"]},
{fishbase: [], inat: ["Kanekonia leichhardti"]},
{fishbase: [], inat: ["Hoplichthys mimaseanus"]},
{fishbase: [], inat: ["Doryrhamphus excisus"]},
{fishbase: [], inat: ["Doryrhamphus melanopleura"]},
{fishbase: [], inat: ["Phyllopteryx dewysea"]},
{fishbase: [], inat: ["Halicampus ensenadae"]},
{fishbase: [], inat: ["Oncorhynchus formosanus"]},
{fishbase: [], inat: ["Oncorhynchus masou"]},
{fishbase: [], inat: ["Oncorhynchus rastrosus"]},
{fishbase: [], inat: ["Oncorhynchus ishikawai"]},
{fishbase: [], inat: ["Salvelinus alpinus"]},
{fishbase: [], inat: ["Salvelinus leucomaenis"]},
{fishbase: [], inat: ["Coregonus hubbsi"]},
{fishbase: [], inat: ["Pseudoxiphophorus anzuetoi"]},
{fishbase: [], inat: ["Pseudoxiphophorus jonesii"]},
{fishbase: [], inat: ["Allodontichtys hubbsi"]},
{fishbase: [], inat: ["Allodontichtys polylepis"]},
{fishbase: [], inat: ["Allodontichtys tamazulae"]},
{fishbase: [], inat: ["Ariosoma hemiaspidus"]},
{fishbase: [], inat: ["Chaetostoma anomala"]},
{fishbase: [], inat: ["Sciades felis"]},
{fishbase: [], inat: ["Sciades leptaspis"]},
{fishbase: [], inat: ["Sciades seemani"]},
{fishbase: [], inat: ["Fragilaria construens binodis"]},
{fishbase: [], inat: ["Chiloglanis devosi"]},
{fishbase: [], inat: ["Chiloglanis kerioensis"]},
{fishbase: [], inat: ["Pinnularia braunii amphicephala"]},
{fishbase: [], inat: ["Olyra taquara"]},
{fishbase: [], inat: ["Oreoglanis hponkanensis"]},
{fishbase: [], inat: ["Engyprosopon osculum"]},
{fishbase: [], inat: ["Monolene maculipina"]},
{fishbase: [], inat: ["Etropus delsmani"]},
{fishbase: [], inat: ["Brachirus breviceps"]},
{fishbase: [], inat: ["Brachirus fitzroiensis"]},
{fishbase: [], inat: ["Pardachirus rautheri"]},
{fishbase: [], inat: ["Pseudaesopia callizona"]},
{fishbase: [], inat: ["Symphurus sitgmosus"]},
{fishbase: [], inat: ["Paraplagusia bleekeri"]},
{fishbase: [], inat: ["Clupea pallasii"]},
{fishbase: [], inat: ["Alosa caspia"]},
{fishbase: [], inat: ["Astyanax rutilus"]},
{fishbase: [], inat: ["Gephyrocharax atricaudatus"]},
{fishbase: [], inat: ["Saccoderma falcata"]},
{fishbase: [], inat: ["Pygopristis denticulatus"]},
{fishbase: [], inat: ["Myloplus zorroi"]},
{fishbase: [], inat: ["Leporinus enyae"]},
{fishbase: [], inat: ["Pterodiscus cookei"]},
{fishbase: [], inat: ["Liza ordensis"]},
{fishbase: [], inat: ["Mugil rammelsbergi"]},
{fishbase: [], inat: ["Planiliza subviridis"]},
{fishbase: [], inat: ["Plecoglossus altivelis"]},
{fishbase: [], inat: ["Galaxias arcanus"]},
{fishbase: [], inat: ["Galaxias mungadhan"]},
{fishbase: [], inat: ["Argentina microphylla"]},
{fishbase: [], inat: ["Galaxias aequipinnis"]},
{fishbase: [], inat: ["Galaxias brevissimus"]},
{fishbase: [], inat: ["Galaxias gunaikurnai"]},
{fishbase: [], inat: ["Galaxias lanceolatus"]},
{fishbase: [], inat: ["Galaxias longifundus"]},
{fishbase: [], inat: ["Galaxias mcdowalli"]},
{fishbase: [], inat: ["Galaxias oliros"]},
{fishbase: [], inat: ["Galaxias supremus"]},
{fishbase: [], inat: ["Galaxias tantangara"]},
{fishbase: [], inat: ["Galaxias terenasus"]},
{fishbase: [], inat: ["Mallotus philippinensis"]},
{fishbase: [], inat: ["Argentina tapetodes"]},
{fishbase: [], inat: ["Monacoa niger"]},
{fishbase: [], inat: ["Nansenia boreacrassicauda"]},
{fishbase: [], inat: ["Dolicholagus longirosytis"]},
{fishbase: [], inat: ["Platybelone argalus"]},
{fishbase: [], inat: ["Tylosurus acus"]},
{fishbase: [], inat: ["Strongylura notata"]},
{fishbase: [], inat: ["Hyporhamphus roberti"]},
{fishbase: [], inat: ["Cheilopogon pinnatibarbatus"]},
{fishbase: [], inat: ["Menidia alchichica"]},
{fishbase: [], inat: ["Menidia ferdebueni"]},
{fishbase: [], inat: ["Menidia labarcae"]},
{fishbase: [], inat: ["Menidia letholepis"]},
{fishbase: [], inat: ["Menidia promelas"]},
{fishbase: [], inat: ["Menidia squamata"]},
{fishbase: [], inat: ["Menidia bartoni"]},
{fishbase: [], inat: ["Menidia charari"]},
{fishbase: [], inat: ["Menidia riojai"]},
{fishbase: [], inat: ["Atherinella pellosemion"]},
{fishbase: [], inat: ["Menidia aculeatum"]},
{fishbase: [], inat: ["Atherinosoma elongatum"]},
{fishbase: [], inat: ["Austromenidia vegia"]},
{fishbase: [], inat: ["Patagonia peregrinum"]},
{fishbase: [], inat: ["Pseudomugil luminatus"]},
{fishbase: [], inat: ["Porophryne erythrodactylus"]},
{fishbase: [], inat: ["Kuiterichthys pietschi"]},
{fishbase: [], inat: ["Antennarius steffifer"]},
{fishbase: [], inat: ["Histrio acuminatus"]},
{fishbase: [], inat: ["Chaunacops spinosus"]},
{fishbase: [], inat: ["Lophiodon spilurus"]},
{fishbase: [], inat: ["Pegasus tetrabelos"]},
{fishbase: [], inat: ["Spinachia linnei"]},
{fishbase: [], inat: ["Trachelochismus aestuarium"]},
{fishbase: [], inat: ["Gouania lineata"]},
{fishbase: [], inat: ["Nettorhamphos radula"]},
{fishbase: [], inat: ["Dellichthys trnskii"]},
{fishbase: [], inat: ["Gadus chalcogramma"]},
{fishbase: [], inat: ["Hymenocephalus iwamotoi"]},
{fishbase: [], inat: ["Trachyrinchus murray"]},
{fishbase: [], inat: ["Trachyrinchus trachyrhynchus"]},
{fishbase: [], inat: ["Trachyrinchus murrayi"]},
{fishbase: [], inat: ["Merluccius gayi"]}
]
You can help by:
(1) checking the 'Explicit deviations for Fishbase recommended by maractwin et al'. Do these make sense? Should any be added or removed?
(2) checking the 'Mystery Synonyms'. These are all not in Fishbase but I wasn't able to identify them as synonyms. They probably need to be manually checked. Some can be swapped into valid fish names. Some should become true 'explicit discrepancies' (new species not yet in Fishbase etc.) and some are probably not even fish (plants etc accidentally grafted to the fish tree).
I've now marked the 6 fish classes as 'complete' down to species meaning no one else except the 'taxon curator(s)' (currently me unless anyone else wants to help) can edit the taxonomy. The idea is that instead of just making taxonomy changes, we should rather propose 'explicit deviations' from Fishbase here and then they can be added to the script which keeps iNat and Fishbase in sync. Its kind of fun to check out the 'trends' tab here: https://www.inaturalist.org/taxa/47178-Actinopterygii and see new fish that are being 'discovered' every few days.
Hopefully, it will be pretty easy to come to consensus about when to 'explicitly deviate' from Fishbase and when not to through discussions here. But just in case there are any disagreements down the road, I propose a working group of top IDers to resolve any disputes. For example, here's the top rayfinned fish identifiers (top 5 globally and top 3 from each continent) which has been proposed as a working group for other groups like odonata:
@maractwin, @sascha_schulz, @richardling, @pmk00001, @ralfmagee, @spela, @jujurenoult, @v_s_, @jpsilva, @nakarb, @lovelyclemmy, @gonodactylus, @davidr, @ianbanks, @johnturnbull, @quiggifur, @fishesoftexas, @fdouglasmartin, @nakarb, @hix, @oryzias, @anudibranchmom, @michaelbakkerpaiva, @stanvrem

December 27, 2017
Tallying observed Vertebrate species with "complete taxa"

As a community, we've observed 1/3 of all known vertebrate species on iNaturalist! Plus, we're ticking an additional 5 to 10 new species each day. How do we know this? We're introducing new functionality that allows us to mark a taxonomic clade on iNaturalist as "complete" and then keep track of how many species on that list are ticked off with observations over time. Below I'll explain how we're using "complete taxa" tools to tally vertebrate species.
Marking a clade as complete means that all extant species in the clade are included in the iNaturalist taxonomy - no more and no fewer. Since vertebrates are marked as complete, you'll now notice new decorations under the taxonomy tab indicating this on taxon pages that fall within it. There is also a section listing "Taxon Curators" who are now the only curators with permissions to edit members of a complete clade.
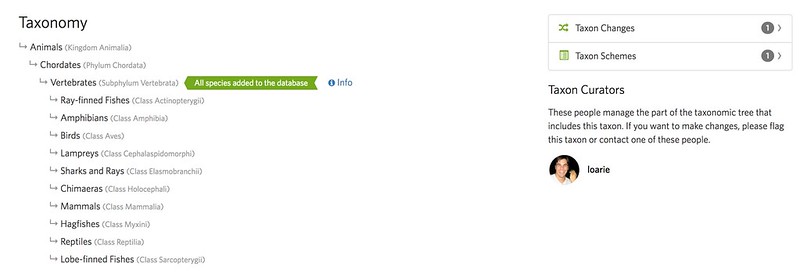
Because complete taxa give us a clearer understanding of all the potential species in a clade, they allow us to track some interesting stats about progress the iNaturalist community has made in ticking these off of the list. At the top of a taxon page that falls within a complete taxon, you'll see a new "Total Species Observed" stat which, at the time of this writing, is "22,538 of 68,295" for vertebrates. The denominator is defined by the "complete" iNaturalist taxonomy and the numerator counts the subset of these species that have been observed. It's kind of remarkable that we've collectively observed 1/3 of all vertebrate species. Considering the thousands of rare deep-sea fishes, tropical herps with tiny ranges, and nocturnal bats and rodents that this number includes, this is a pretty amazing accomplishment!.
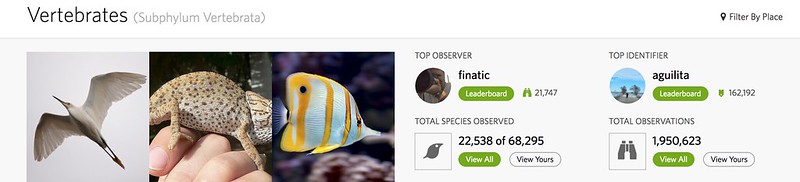
Under the trends tab, below the "Trending" feed there are two new feeds. The first is "Discoveries." This shows the most recent newly-identified species in the taxon. The second is "Wanted" which shows species in the taxon that have not yet been observed.
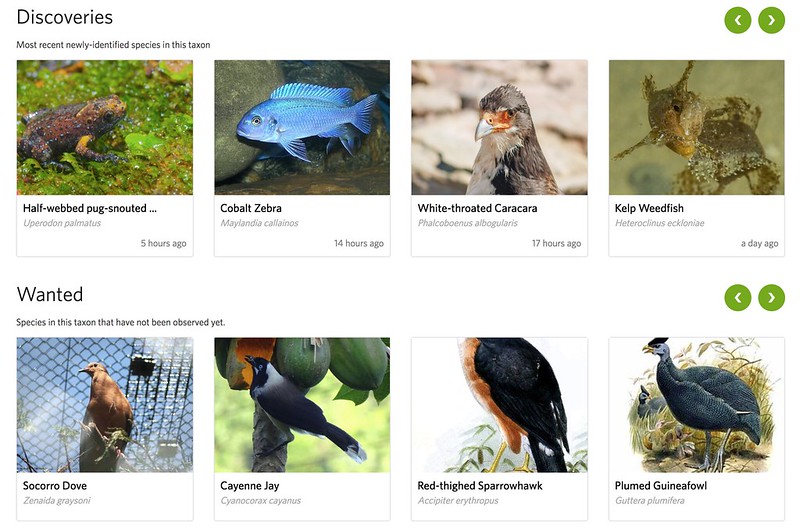
I did a quick analysis of the vertebrate discoveries from the last 38 days and I count:
3 other fishes
Here's how those discoveries over the last 38 days stack up over time. It's pretty exciting to see that we're getting about 5-10 new vertebrates added to iNaturalist each day. At that rate it will take us about 4 more years to reach half of all vertebrates. It will be interesting to see if that rate gets damped by increasingly hard-to-get species or boosted by a growing iNaturalist community.
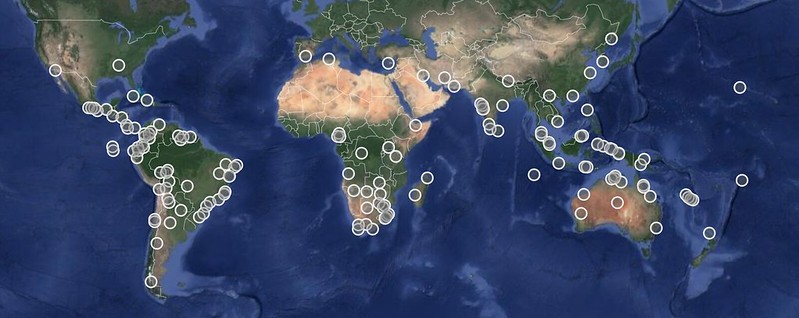
You can click through to see the actual observations responsible for the growth in Discoveries. Mapping them reveals that most of these discoveries are coming from the tropics of the world. For example,
here's a map of the bird, mammal, reptile, and amphibian discoveries from the last 38 days. This should come as no surprise since the tropics harbor the richest biodiversity on the planet and also represent some of the least-explored areas by the iNaturalist global community.
Two caveats: first, the discovery feed pulls from verifiable observations. Some are the result of mis-IDs. But they disappear from the feed as they get corrected. Second, some of the entries in the discoveries feed are triggered by taxonomic changes. For example, if the scientific name of a species changes, then the first identification of the output species will trigger an entry in the feed even though there were already identifications of the input taxon. We're working on a solution to keep these out of the feed, but in the meantime I tried to manually exclude these cases from the numbers above, but there are less than 5 complex cases that remain (e.g. when a species was in the system as two active synonyms representing the same taxa and then swapped together).
Taxonomic Details
Taxonomy is still very much a work in progress. It may come as surprise to some that we don't yet have a global list for plants or even butterflies. This means that the taxonomy for many branches of the tree of life for iNaturalist will inevitably be messy. Some taxa will be missing, while others will be duplicated as "synonyms" (different names that all refer to the same taxon). Fortunately, the branch of life representing we humans and our closest relatives, the vertebrates, is relatively well-known. For each of the 10 classes making up this subphylum, there are relatively well-maintained and continually updating checklists that list all extant species members globally. iNaturalist has adopted several of these lists as external taxonomic references. They are: FishBase for the 6 fish Classes, Amphibian Species of the World for amphibians, the Reptiles Database for reptiles, the Clements Checklist of Birds of the World for birds, and the IUCN RedList itself for mammals.
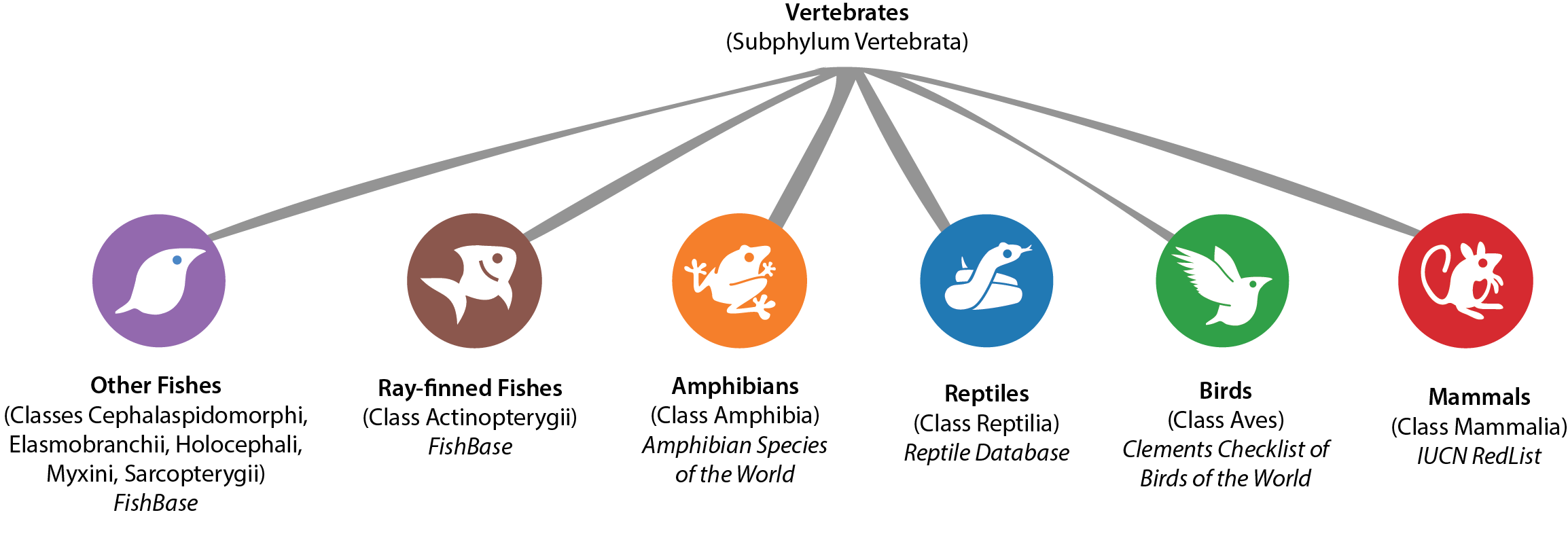
While new species of vertebrates are being described each day, these external taxonomic references at least strive to be relatively complete and to keep relatively up to date. Keeping iNaturalist in sync with these lists should, in theory, reduce curation work and taxonomic confusion for everybody
However, it's difficult to get the entire iNaturalist community onboard with a single external taxonomic reference. The inevitable errors, lag-time in adopting new species, and subjective taxonomic opinions in any list will frustrate some members of the iNaturalist community. When this happens, we can make an exception against using the external taxonomic reference for the controversial taxon/taxa in question by making an explicit discrepancy. For example, the Reptile Database considers just one species of Mountain Kingsnake in California, Lampropeltis zonata sensu lato (meaning in the broad sense). However, the most widely used regional reptile reference in California considers these snakes to represent 2 species, Lampropeltis multifasciata and Lampropeltis zonata sensu stricto (meaning in the narrow sense). It might therefore make sense for iNaturalist to make an explicit discrepancy to avoid following the Reptile Database's decision to merge Lampropeltis multifasciata & Lampropeltis zonata sensu stricto into Lampropeltis zonata sensu lato.
I've created journal posts to serve as discussions for proposing and making decisions about explicit discrepancies for
reptiles, fishes, mammals, and amphibians. Please contribute to those conversations if you'd like to discuss explicit discrepancies for each of these clades.

December 28, 2017
Explicit Discrepancies from Clements for Birds?
@maxkirsch and I have been working to keep iNaturalist in sync with Clements for birds. Motivated by this post here's a place to propose/discuss any Explicit Discrepancies from the Clements Checklist of Birds of the World.
By explicit discrepancies I mean something like this fake example:
"Clements splits Troglodytes troglodytes (sensu lato) into Troglodytes pacificus ,Troglodytes hiemalis, and Troglodytes troglodytes (sensu stricto). My understanding is that most ornithologists no longer believe that there are differences between these birds, I propose we explicitly deviate from Clements by lumping these as Troglodytes troglodytes (sensu lato)"
This is not a place to propose switching to the IOC World Bird List. But if it turns out that the difference between (Clements + our explicit discrepancies) is closer to IOC than Clements that might be something to consider.
Here's the analysis of top global and regional bird IDers I've been making for various groups of vertebrates. They are:
@markuslilje, @ivanresendizcruz, @d_kluza, @john8, @greglasley, @joshuagsmith, @psweet, @kokhuitan, @momoto-erick, @paultavares, @reddad, @roby, @martingrimm, @inasiebert, @johnnybirder, @jakob, @gawie, @jadonald, @adammyates, @d_kurek, @dpom, @hfabian, @gyrrlfalcon, @cliygh-and-mia, @scops, @amarzee, @camilojotage, @nelson_wisnik, @tegorl
Thanks!
Scott
Archives
- Month Posts
- November 2023 1
- February 2023 1
- January 2023 5
- November 2022 2
- October 2022 1
- May 2022 1
- February 2022 1
- August 2021 1
- July 2021 1
- April 2021 1
- July 2019 3
- May 2019 1
- April 2019 3
- January 2019 2
- August 2018 3
- July 2018 2
- May 2018 2
- January 2018 1
- December 2017 3
- October 2017 2
- September 2017 2
- August 2017 1
- May 2017 1
- April 2017 1
- December 2016 1
- July 2016 1
- March 2016 1
- February 2016 4
- January 2016 1
- November 2015 1
- October 2015 1
- April 2015 1
- March 2015 1
- August 2014 1
- July 2014 2
- March 2014 1
- September 2013 1
- May 2013 2
- April 2013 1
- March 2013 1
- February 2013 2
- September 2012 1
- March 2012 1
- February 2012 1
- December 2011 1
- October 2010 1


